Cutting up the genes – the structure of a spliceosome
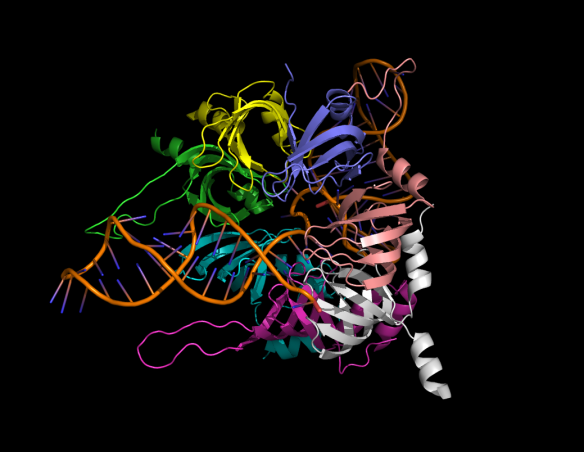
What does it look like?
Cartoon representation of the core subunits of the U4 small nucleolar ribonuceoprotein complex (U4 snRNP). The seven protein core subunits of the complex are shown in ribbon representation SmE-SmG-SmD3-SmB-SmD1-SmD2-SmF. The proteins form a ring structure encircling the core RNA part of the complex.
What is it?
Most of our genes are made up of both introns and exons. While exons encode proteins, introns must be removed to produce a functional messenger RNA for protein production by the ribosome. Intron removal (called splicing) is catalyzed by a large and dynamic complex called the spliceosome, which contains a number of small nuclear ribonucleoproteins (snRNPs – pronounced 'snurps'). Most snRNPs contain seven Sm proteins (SmB/B', SmD1, SmD2, SmD3, SmE, SmF and SmG) in common. The structure above shows how these seven Sm proteins assemble into a beautiful ring structure, that encircles the RNA component of snRNPs to provide stability and protection. The ability of researchers to capture such large and dynamic biomolecular structures in a crystalline state never ceases to amaze.
Where did the structure come from?
This structure was solved in the lab of Kyoshi Nagai at the Laboratory for Molecular Biology in Cambridge (UK). You can view it at http://www.rcsb.org/pdb/explore/explore.do?structureId=2Y9A and read more about it in http://www.ncbi.nlm.nih.gov/pubmed/21516107.