Pass the Electron – The First Membrane Protein Structure Shows How Purple Bacteria Get Energy from Light
What does it look like?
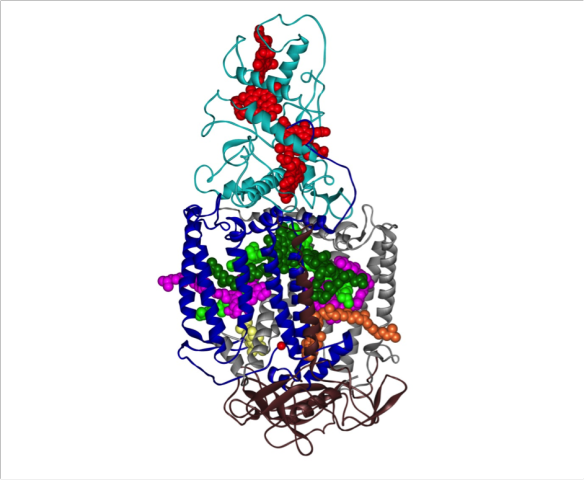
The crystal structure of the photosynthetic reaction centre from the purple bacterium Rhodopseudomonas viridis.
What is it?
In 1985 Johann Deisenhofer and colleagues successfully solved the first membrane protein structure to atomic resolution by X-ray crystallography [1]. The protein complex shown is the photosynthetic reaction centre from the purple bacterium Rhodopseudomonas viridis and for solving this structure Johann Deisenhofer and Hartmut Michel won the 1988 Nobel Prize in Chemistry [2]. Some bacteria, like plants, can perform photosynthesis, the process of acquiring energy from light. The photosynthetic reaction centre is the membrane protein complex in the R. viridis membrane that captures the light energy needed for photosynthesis to occur and converts it to chemical energy.
The photosynthetic reaction centre consists of four protein subunits called heavy (brown ribbons), medium (blue ribbons), light (grey ribbons) and cytochrome (cyan ribbons). The long helices in the heavy and medium subunits are the parts of the protein that reside in the core of the cell membrane. There are also a number of co-factors: four bacterio-chlorophyll molecules (light and dark green spheres), two bacterio-pheophytins (pink spheres), one non-haem iron atom (single red ball), two quinones (QA (orange spheres) and QB (yellow spheres)) and four haem groups (red spheres). When a photon of light is absorbed by a pair of bacterio-chlorophylls, the pair goes into an excited state and loses an electron which is passed to the bacterio-pheophytins in the super quick time of 3 pico-seconds. The electron is then passed to QA and then to QB via the non-haem iron. At this point the QB will leave the complex and establish a gradient of protons across the bacterial membrane which will then be used by other proteins to generate the cell's energy currency, a molecule called ATP. The pair of bacterio-chlorophyll at the start of this cycle gets replenished with electrons from the haem groups in the cytochrome domain so that they return to the ground state from their excited state. The system is now reset and the whole cycle of passing electrons around can start again with the next photon of light.
Where does it come from?
The above structure was solved by J. Deisenhofer et al. and is from the protein data bank (PDB number 1PRC [3]) and the following references:
[1] J. Deisenhofer, O. Epp, K. Miki, R. Huber, H. Michel: Structure of the protein subunits in the photosynthetic reaction centre of Rhodopseudomonas viridis at 3 Å resolution. Nature 1985, 318: 618-624.
[2] J. Deisenhofer, H. Michel: Nobel Lecture: The photosynthetic reaction centre from the purple bacterium Rhodopseudomonas viridis. EMBO J. 1989, 8: 2149-2170.
[3] J. Deisenhofer, O. Epp, I. Sinning, H. Michel: Crystallographic refinement at 2.3 Å resolution and refined model of the photosynthetic reaction centre from Rhodopseudomonas viridis. J. Mol. Biol. 1992, 246: 429-457.